 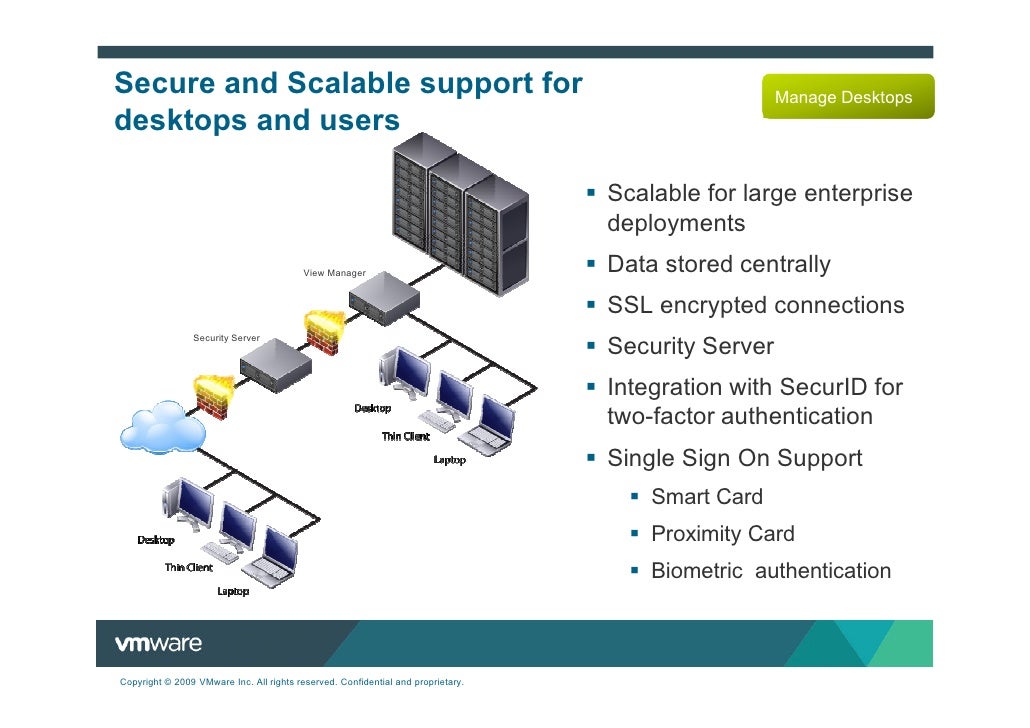
NetworkView is implemented as a plugin to the molecular visualization software VMD. network analysis of the complex to explore the allosteric network in the protein and monobody. 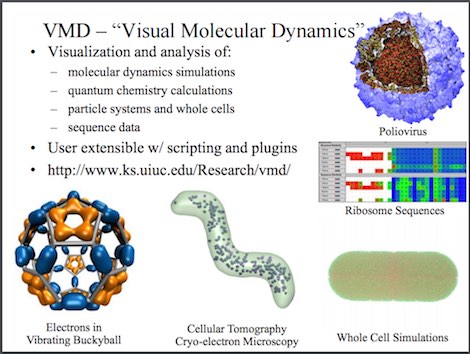
An API reference for NetworkView has been generated as Tcldoc documentation. Residue-Residue Contact Changes during Functional Processes Define Allosteric Communication Pathways. One of the plugin of VMD, NetworkView 56, was used for the topological. Luthey-Schulten, NetworkView: 3D display and analysis of. NetworkView projects the network onto the underlying 3D molecular structure so that visualization and analysis of the network can be coupled to physical and biological properties. For a general introduction on how to prepare and analyze networks using NetworkView, visit the NetworkView Tutorialpage. Using the popular visualization program visual molecular dynamics (VMD). These networks typically model individual protein residues and nucleic acid monomers as nodes and their pairwise contacts as edges with associated weights. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
